Introduction by Example
We shortly introduce usage basics of AMPy through several self-contained examples. Up to this momemnt, you can follow this tutorial in the format of Colab Notebook or check out attached video (TBD).
Essentially, AMPy allows you to perform the following processing procedures:
Extract Kinematics from Raw Video
We will show a simple example of extracting trajectories from the predefined video:
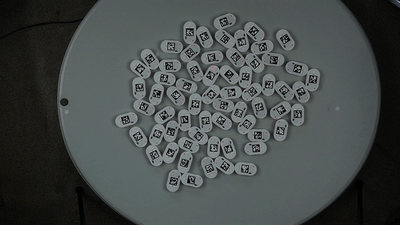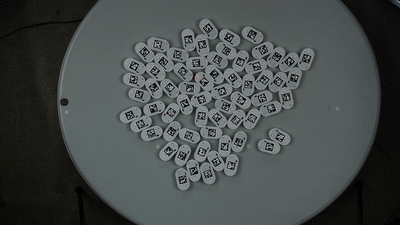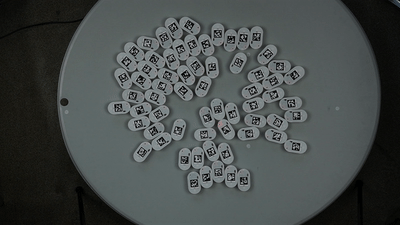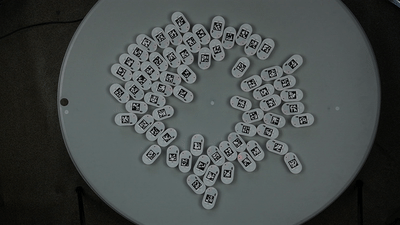
To extract the robots’ trajectories from the video, we import the Processor class from ampy.processing and create the corresponding object:
from ampy.processing import Processor
VP = Processor()
Then we pass the videofile path using set_filename method:
filename = 'test_video.mp4'
VP.set_filename(filename)
From this moment we can extract system’s Cartesian kinematics by the cartesian_kinamatics function:
cart_kin = VP.cartesian_kinematics(bots_number=65,
begin_frame=120,
end_frame=1800,
get_each=5,
ignore_codes=(),
scale_parameters=(1, 0))
We can see that this method holds 6 parameters: bots_number is a number of tracking objects presented in the video; begin_frame and end_frame describe a start/stop frames for kinematics extraction; get_each sets frames decimation frequency (to speed up the execution); ignore_codes is a list of markers’ ids which are not considered during the tracking; scale_parameters correspond to the α and β parameters of a frame linear transformation (adjustable contrast and brightness parameters).
To extract the polar representation of kinematics, you should provide the coordinates of the field center. This can be done automatically using field_center_auto if you place additional markers on the area’s borders. Otherwise, we can set it up manually:
center = (1920 // 2, 1080 // 2)
polar_kin = VP.polar_kinematics(cartesian_kinematics=cart_kin,
field_center=center)
In some cases, it can be beneficial to convert linear distances from pixels to centimeters. Scaling factor of such transformation can be obtained via metric_constant with respect to the size of ArUco markers:
marker_size = 3 # in centimeters
scaling_factor = VP.metric_constant(marker_size=marker_size, scale_parameters=(1, 0))
Note
If you are lucky to have your own tracking software, you can still use AMPy to evaluate various statistical characteristics. In order to do that, it is required to convert your data to the following format (per frame): [object_id, orientation_angle, object_center_coordinate].
Evaluate Group Dynamics
Module statistics2d allows you to evaluate several characeristics represented in the form of temporal dependencies.
The one can obtain mean displaiments of robots from the center by the means of the
mean_distance_from_centerfunction:
from ampy.statistics2d import mean_distance_from_center
distance = mean_distance_from_center(kinematics=polar_kin)
Mean polar angle of robots in the system can be calculated via
mean_polar_angle:
from ampy.statistics2d import mean_polar_angle
angle = mean_polar_angle(kinematics=polar_kin)
On top of that, you can evaluate mean polar angle in sense of the angular path using
mean_polar_angle_absolute:
from ampy.statistics2d import mean_polar_angle_absolute
angle_abs = mean_polar_angle_absolute(kinematics=polar_kin)
Mean squared distance (to the center of the field) can be evaluated by the
mean_cartesian_displacementsfunction:
from ampy.statistics2d import mean_cartesian_displacements
cart_disp = mean_cartesian_displacements(kinematics=cart_kin)
In order to check whether system’s configuration corresponds to some regular lattice, you can apply
bond_orientationwith the order parameterneighbours_number:
from ampy.statistics2d import bond_orientation
boo = bond_orientation(kinematics=cart_kin, neighbours_number=6, folds_number=6)
Spatio-temporal correlation of the system can be evaluated by the
chi_4function:
from ampy.statistics2d import chi_4
from multiprocessing import Pool
import os
data = []
for time in time:
data.append((cart_kin, time, 100))
with Pool(os.cpu_count()) as pool:
stcp = pool.starmap(chi_4, data)
Clustering coefficient of the system can be obtained by the
cluster_dynamicsfunction:
from ampy.statistics2d import cluster_dynamics
cl_coeff = cluster_dynamics(kinematics=cart_kin)
This function has an optional parameter collide_function specifying collision rules for robots.
Correlations between robots positions, orientations and velocities can be evaluated by the following functions:
position_correlation,orientation_corrilation, andvelocity_correlation. For simplicity, we will evaluate them in the 400x400 window:
from ampy.statistics3d import position_correlation, orientation_correlation, velocity_correlation
pos_corr = position_correlation(kinematics=cart_kin, x_size=200, y_size=200)
orient_corr = orientation_correlation(kinematics=cart_kin, x_size=200, y_size=200)
vel_corr = velocity_correlation(kinematics=cart_kin, x_size=200, y_size=200)
To provide better visual summary, you may average correlation maps for all processed frames:
pos_corr = np.mean(np.array(pos_corr), axis=0)
orient_corr = np.mean(np.array(orient_corr), axis=0)
vel_corr = np.mean(np.array(vel_corr), axis=0)